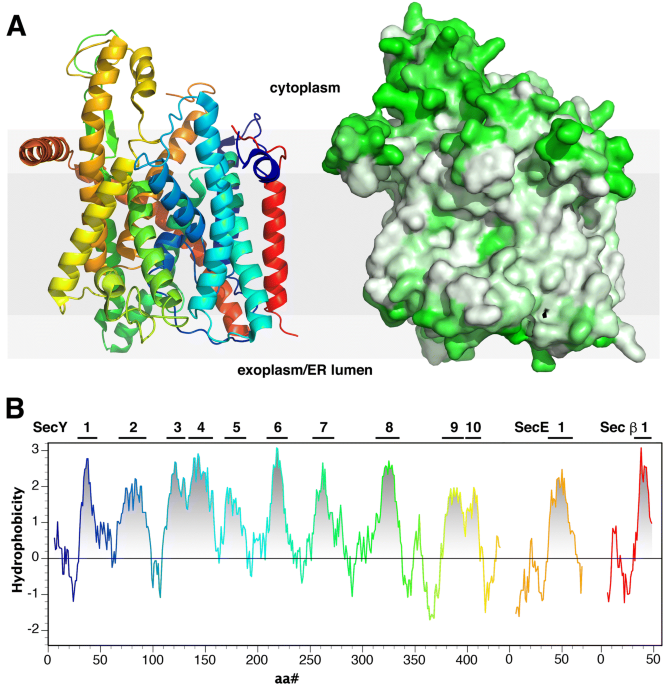
Side-chain hydrophobicity scale derived from transmembrane protein folding into lipid bilayers

Side-chain hydrophobicity scale derived from transmembrane protein folding into lipid bilayers
Membrane Protein Integration and Topogenesis at the ER

Lipid bilayer regulation of membrane protein function: gramicidin channels as molecular force probes
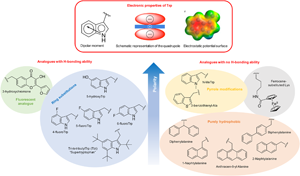
Tryptophan, more than just an interfacial amino acid in the membrane activity of cationic cell-penetrating and antimicrobial peptides, Quarterly Reviews of Biophysics
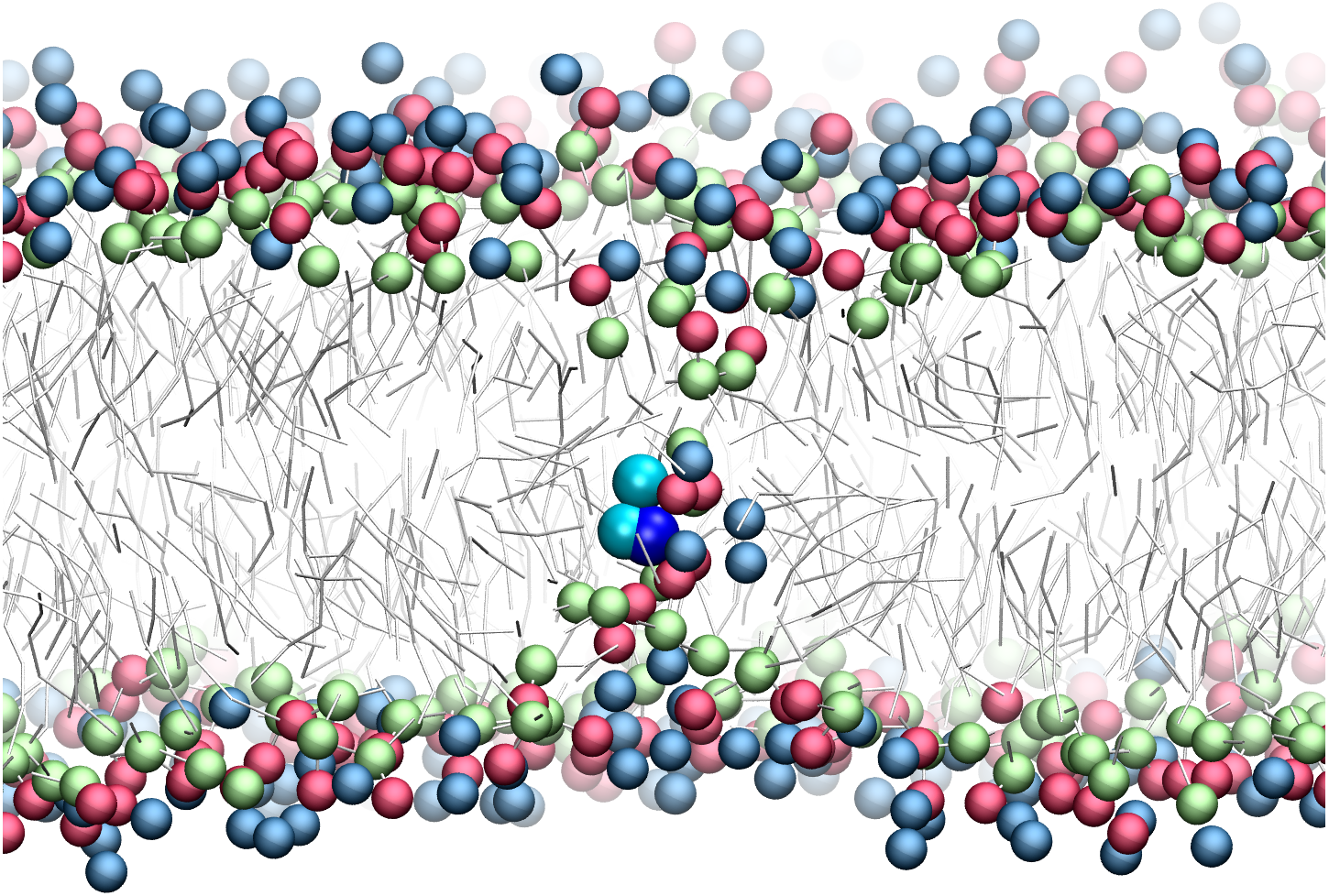
Refining amino acid hydrophobicity for dynamics simulation of membrane proteins [PeerJ]

The Hydrophobic Effect Is a Principal Force Stabilizing Tertiary and Quaternary Structures - LabXchange

How bilayer properties influence membrane protein folding - Corin - 2020 - Protein Science - Wiley Online Library
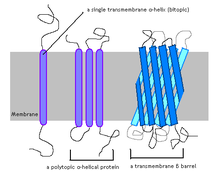
Structural Biochemistry/Chemical Bonding/Hydrophobic interaction/Hydrophobicity scales - Wikibooks, open books for an open world
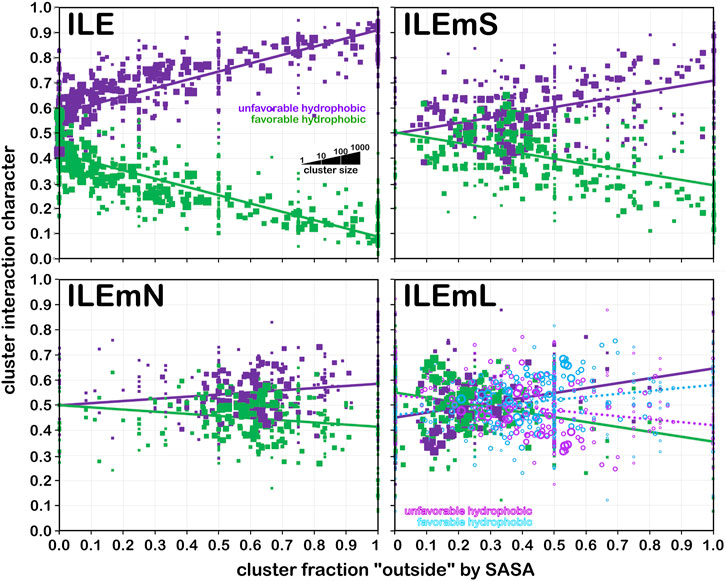
Frontiers 3D interaction homology: The hydrophobic residues alanine, isoleucine, leucine, proline and valine play different structural roles in soluble and membrane proteins
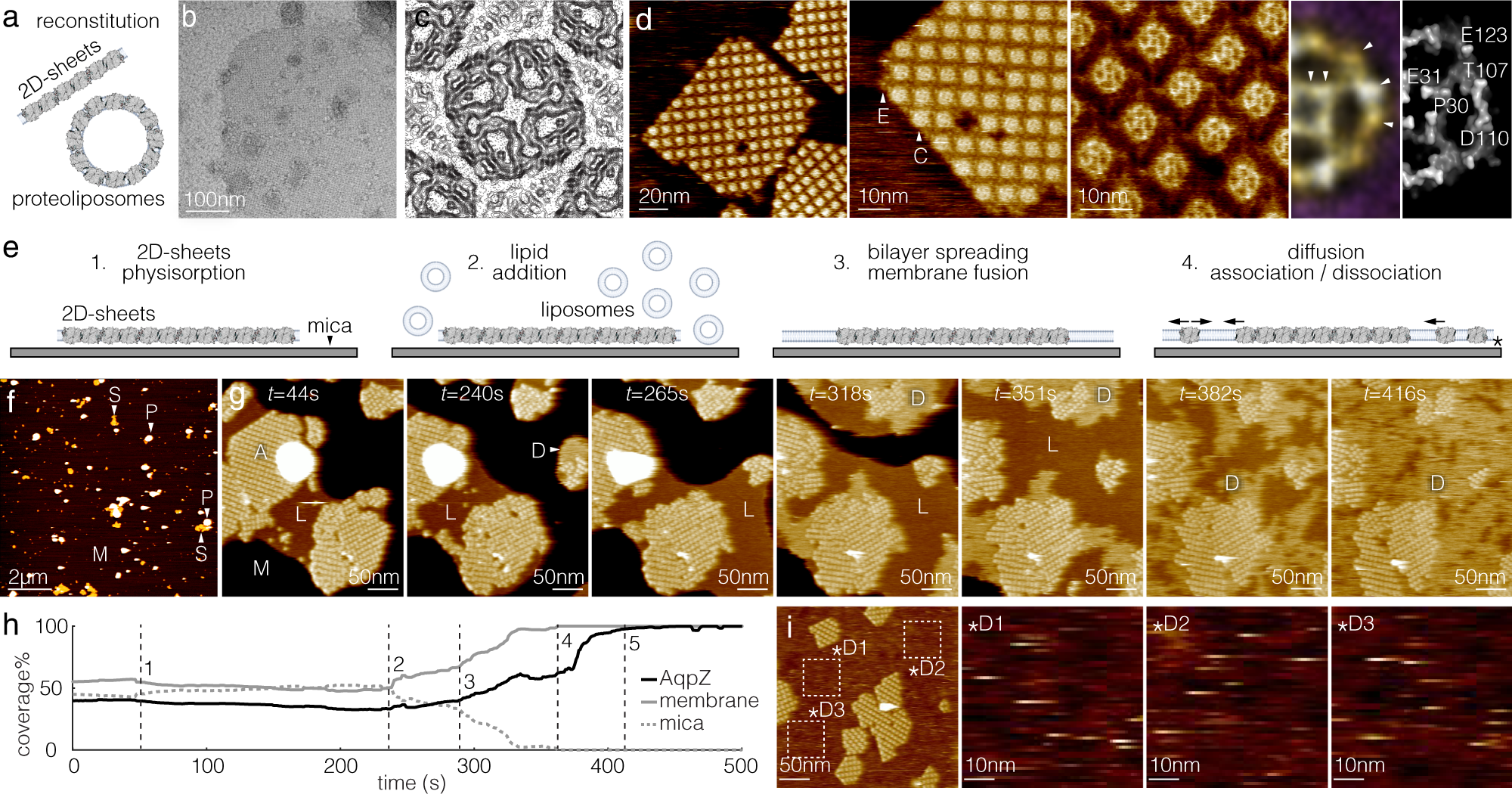
Membrane-mediated protein interactions drive membrane protein organization
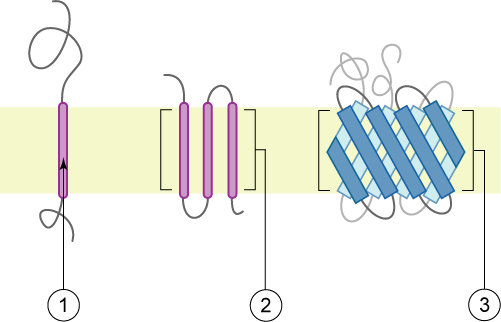
Transmembrane protein - Wikipedia
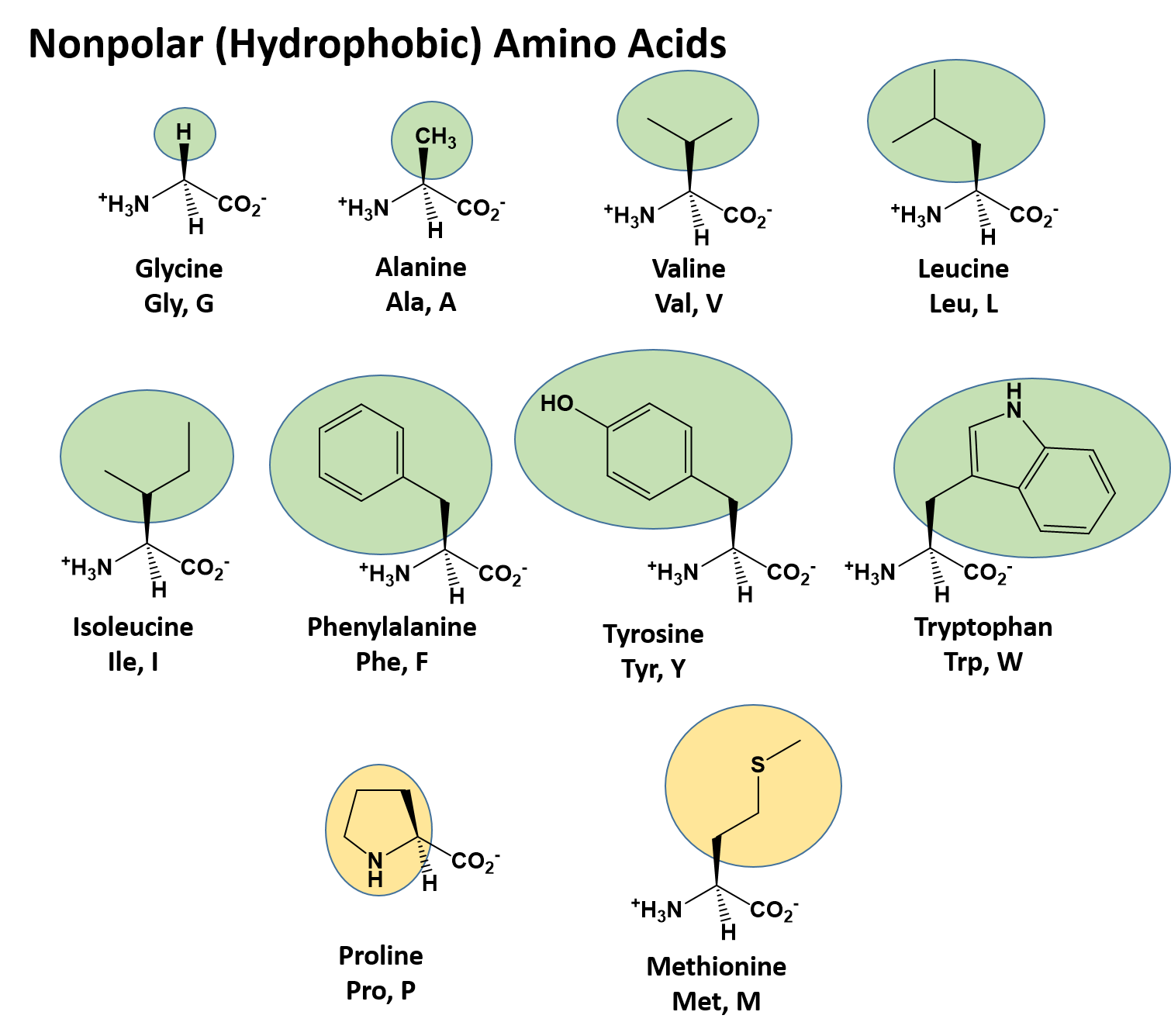
Chapter 2: Protein Structure - Chemistry

Dawn of a New Era for Membrane Protein Design